COPD clustering papers often justify the importance of their clusters by demonstrating their association to a meaningful clinical outcome, such as the frequency of respiratory exacerbations or rate of disease progression. In previous posts, the point has been made that the COPD “phenotypic space” is usually a continuum and that continuous “disease axes” provide a more accurate and reproducible summarization of this continuum than clusters. This post describes one possible response to the challenge of poor cluster reproducibility by identifying disease axes through supervised learning. At the end, a way back to subtypes is explored.
First, if the goal is to be able to make better predictions of future COPD outcomes, it’s clear that for disease axes generally outperform clusters, often by a substantial margin. In a paper from 2018, Greg Kinney and colleagues at National Jewish Health used factor analysis to identify four main factors of variability in COPDGene data, including measures from chest CT, spirometry, and functional assessments. Focusing on overall mortality, each of the four factors was significantly associated to mortality, and there was synergistic effect of factors related to emphysema and airways disease on mortality risk, as shown below.
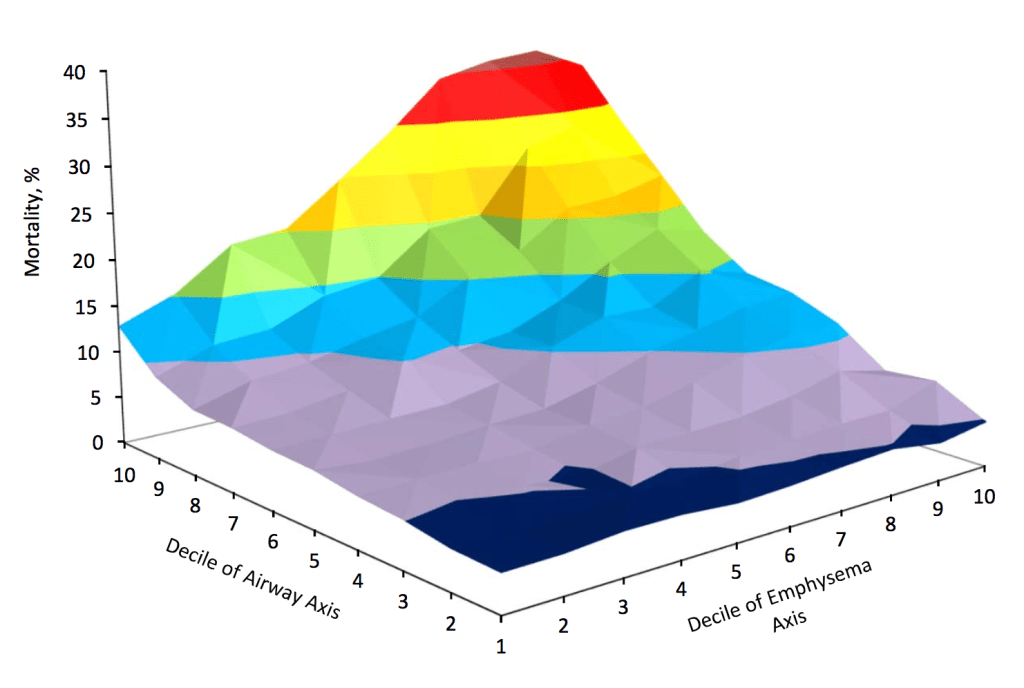
The mortality analysis from Kinney’s paper encourages us to think about defining subtype boundaries based on the distribution of outcome risk is relation to disease axes. Unlike clustering, which defines groups based on similarity in the original clustering space, this approach defines a relevant feature space (defined by disease axes) but then establishes group boundaries based on the local probability of a given outcome. As this paper shows, in the space defined by emphysema and airways disease, the relationship to mortality is non-linear. It would be natural to divide subjects into two groups along the boundary where mortality risk begins to increase sharply. One benefit of this outcome-driven approach to defining subtypes is that it can easily be adapted to other relevant outcomes, for example treatment response.
But what about a head-to-head comparison of outcome prediction for subtypes versus disease axes? In a 2019 paper in Thorax, we used prediction models to generate subtype-oriented disease axes (SODAs). Unlike disease axis from factor analysis or PCA, the use of supervised learning to generate SODAs allows the user more explicit control over what their disease axes mean/represent. In factor analysis and PCA, the orientation of a disease axis is determined by the correlation structure of the original data. While this is often desirable, it means that axes are determined by the initial selection of variables. SODAs on the other hand are determined by:
- the response variable, which in this case is a binary variable encoding two subgroups of subjects (i.e. subtypes)
- input or predictor variables
To make a SODA, we build a predictive model using the two subtype groups (i.e. subtypes) as a binary response and the input variables as predictors. For example, if you specify a group of “pure airways disease” subjects and another group of “emphysema predominant subjects”, then the resulting SODA axis can be thought of as a line with airways disease at one end and emphysema at the other. Importantly, the SODA model can be built with a subset of subjects and then applied to the entire dataset.
Using that method, you can generate SODAs and compare them directly to their “parent” subtypes. In 4,726 subjects from COPDGene with 5-year followup data we generated 6 SODAs from 6 subtype pairs, and we compared the predictive performance of the SODAs to their corresponding subtypes with respect to five-year prospective change in FEV1 and CT emphysema. As shown in the table below, SODAs consistently explained more variance in COPD progression than subtypes. If you put SODAs and subtypes in the same models (Table 2 from the paper), the SODAs were nearly always significant (9 out of 12 models) whereas the subtypes were not (4 out of 12).
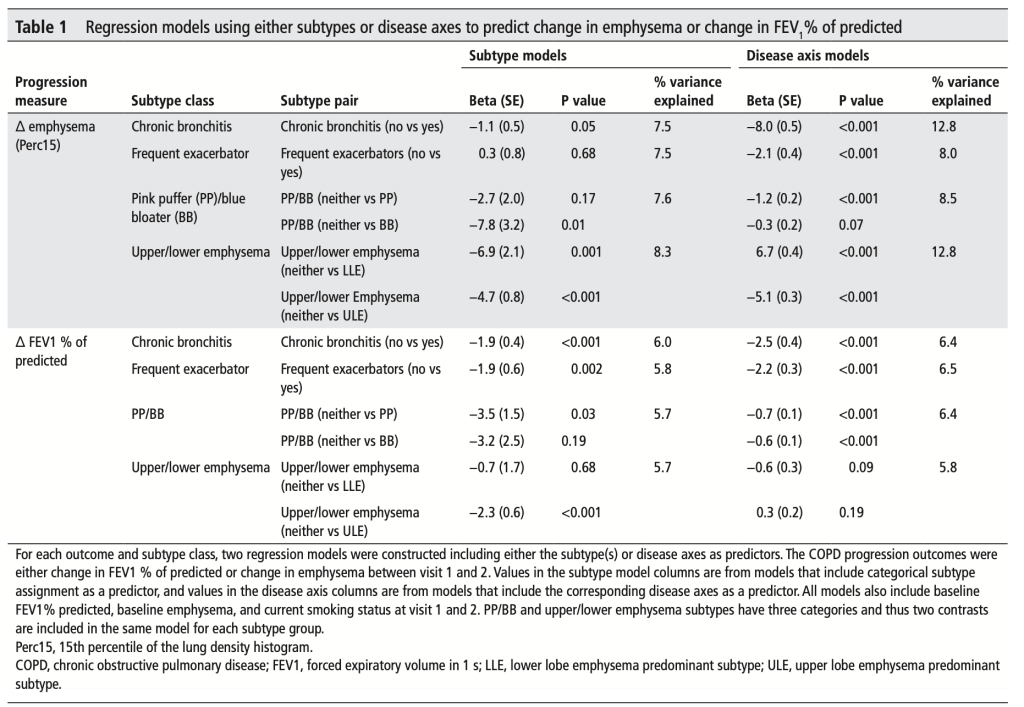
If one focuses only on the subtype of chronic bronchitis (per the ATS-DLD definition), the bronchitis SODA explains nearly twice as much of the 5-year change in emphysema as the original variable. Why is this? First of all, let’s look at how the chronic bronchitis SODA is distributed according to chronic bronchitis status at baseline and at the follow-up visit.
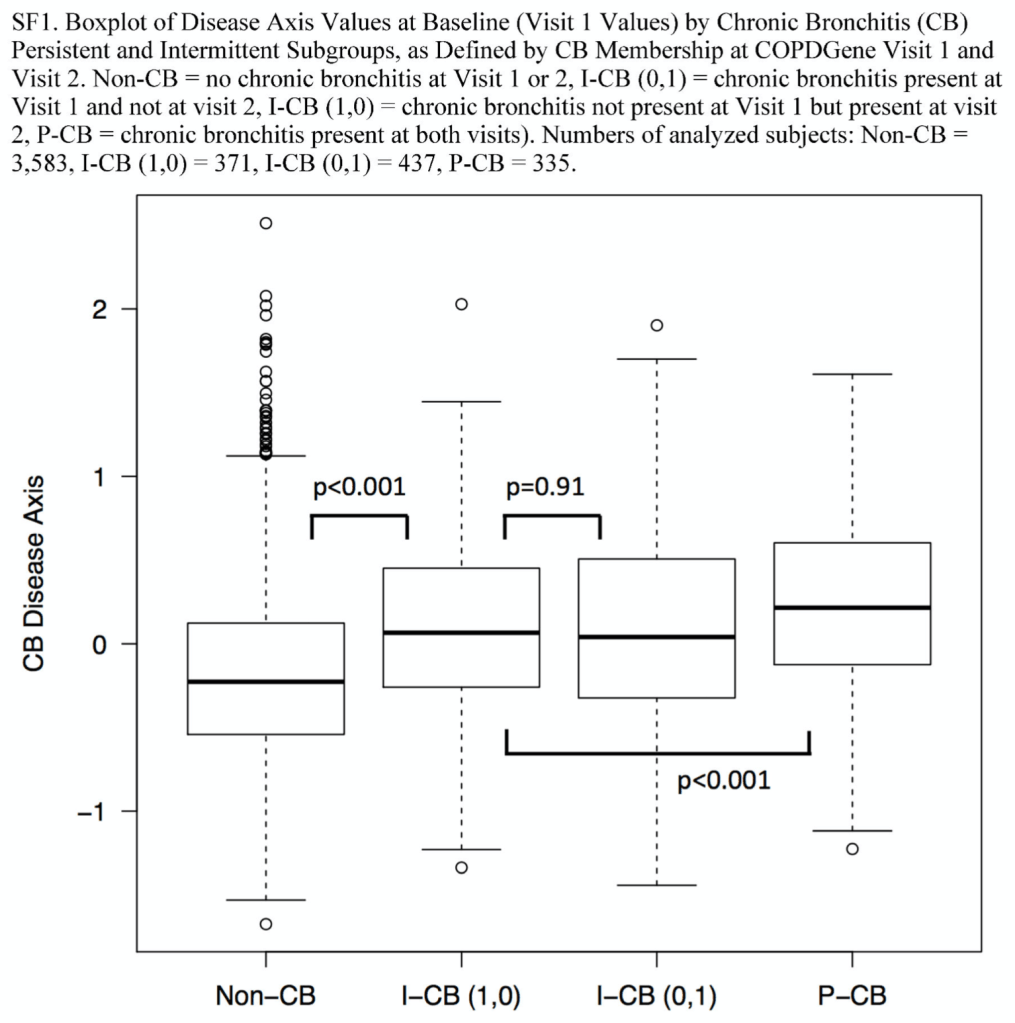
It is evident that the values of the SODA are shifted as one would expect. Subjects with persistent chronic bronchitis (the P-CB group, i.e. chronic bronchitis at baseline and at five-year followup) have higher SODA values than those without chronic bronchitis (the left group) or those with bronchitis at only one visit (the two middle groups). Importantly, the SODA models were built using information only from the baseline visit.
It’s also fun to look at loess curves of the relationship between the bronchitis SODA and prospective five year changes in FEV1 and emphysema. You can see why the emphysema prediction is particularly strong.
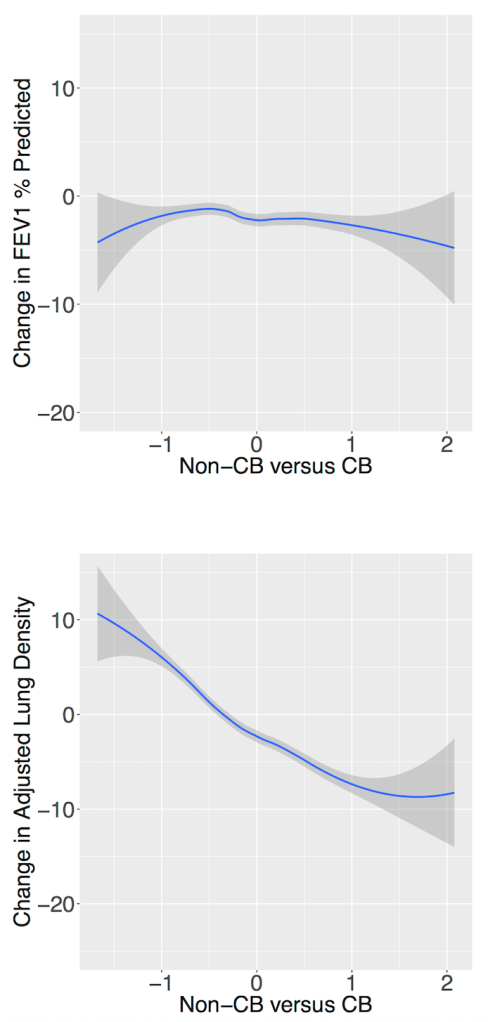
So, how is it that a SODA that is trained by a subtype can actually predict better than the subtype itself? The biggest reason is that the SODA models have access to a lot of information beyond the subtype assignment, namely through their access to the predictor variables. The regression model can be thought of as a filter for the predictor variables. It sorts through the predictors, keeping only those aspects of the variability that are relevant to the subtype. In the case of the chronic bronchitis SODA, it includes not just chronic bronchitis information, but other relevant aspects of FEV1, CT emphysema, and airway hyperresponsiveness. It turns out that, in addition to being relevant to chronic bronchitis, this information is also relevant to future emphysema progression.
So perhaps I’ve convinced you that disease axes are great and the way to go, at least in certain cases. But sometimes you just need subgroups, not axes. For these situations, there is a way to get back to subgroups, and that way is probably more principled and reproducible than clustering. To extend our earlier idea about defining subgroups according to the distribution of outcome risk within a relevant feature space, we can do this for the bronchitis SODA using classification trees, for example. The image below shows subgroups defined solely by cutpoints in the bronchitis SODA, where cutpoint locations were determined by maximizing the difference in CT emphysema progression between groups. The box and whisker plots below show the distribution of change in Perc15 values for each of the bronchitis SODA subgroups. (This analysis was done at an earlier time when there were fewer COPDGene subjects with available follow-up data, hence the smaller numbers.)
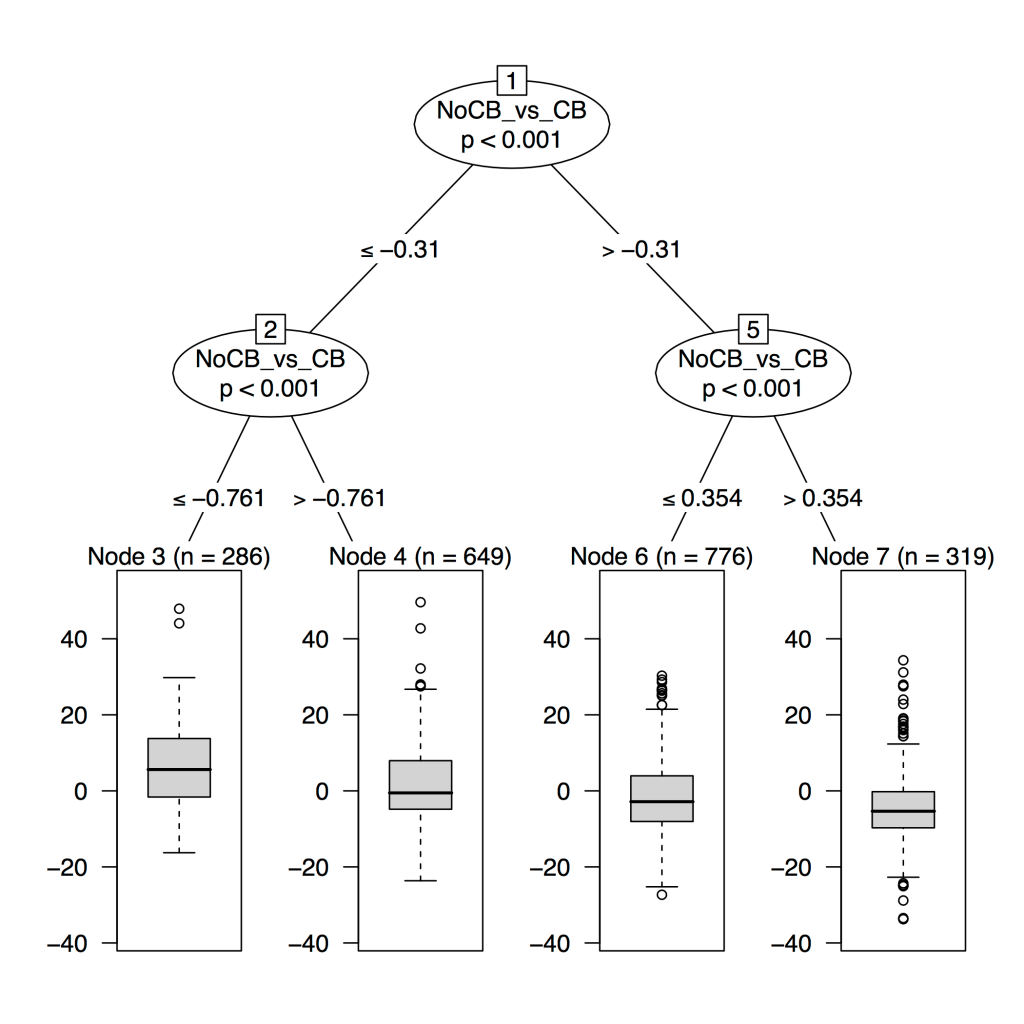
To sum up, this post describes an alternative to clustering that relies heavily on supervised learning to define disaease axes, but it can bring you back to subgroups if you wish, and those subgroups are more likely to be relevant for a given outcome than unsupervised clusters. If we are concerned primarily about the association of our subtypes to some important COPD measure (and we usually are), then it makes sense to incorporate supervised learning early in the subtype identification process to define relevant feature spaces for subsequent cluster/subtype identification.
[…] We should use supervised learning methods rather than focusing exclusively on unsupervised clustering. The ability of supervised models to help identify (or construct) important features can really help us in the first stage of data exploration. Too often, unsupervised clustering yields multiple possible solutions with only modest associations to COPD outcomes and weak justification for choosing a single, “optimal” solution. Supervised methods are much better equipped to discriminate between models and reduce data dimensionality in a way that is relevant to COPD-related outcomes, as discussed here. […]
LikeLike
[…] 4: A clustering alternative: supervised prediction for disease axis discovery. Makes the case for an alternative path to clusters that goes first through supervised prediction and disease axes, then to outcome-driven subtypes. […]
LikeLike
[…] axes that have a very clear and specific meaning. As a bonus, these disease axes often provide more accurate prediction of future COPD events than subtype […]
LikeLike